A | B | C | D | E | F | G | H | CH | I | J | K | L | M | N | O | P | Q | R | S | T | U | V | W | X | Y | Z | 0 | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9

| Part of a series on |
| Genetic genealogy |
|---|
| Concepts |
| Related topics |
In human genetics, a human Y-chromosome DNA haplogroup is a haplogroup defined by mutations in the non-recombining portions of DNA from the male-specific Y chromosome (called Y-DNA). Many people within a haplogroup share similar numbers of short tandem repeats (STRs) and types of mutations called single-nucleotide polymorphisms (SNPs).[2]
The human Y-chromosome accumulates roughly two mutations per generation.[3] Y-DNA haplogroups represent major branches of the Y-chromosome phylogenetic tree that share hundreds or even thousands of mutations unique to each haplogroup.
The Y-chromosomal most recent common ancestor (Y-MRCA, informally known as Y-chromosomal Adam) is the most recent common ancestor (MRCA) from whom all currently living humans are descended patrilineally. Y-chromosomal Adam is estimated to have lived roughly 236,000 years ago in Africa[citation needed]. By examining other bottlenecks most Eurasian men (men from populations outside of Africa) are descended from a man who lived in Africa 69,000 years ago (Haplogroup CT). Southeast Asia has been suggested as the origin for all non-African human Y chromosomes,[4] however, this has cited as being unlikely.[5] Other major bottlenecks occurred about 50,000 and 5,000 years ago and subsequently the ancestry of most Eurasian men can be traced back to four ancestors who lived 50,000 years ago, who were descendants of African (E-M168).[6][7][8]
Naming convention
This section has multiple issues. Please help improve it or discuss these issues on the talk page. (Learn how and when to remove these template messages)
|
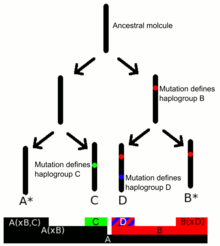
Y-DNA haplogroups are defined by the presence of a series of Y-DNA SNP markers. Subclades are defined by a terminal SNP, the SNP furthest down in the Y-chromosome phylogenetic tree.[9][10] The Y Chromosome Consortium (YCC) developed a system of naming major Y-DNA haplogroups with the capital letters A through T, with further subclades named using numbers and lower case letters (YCC longhand nomenclature). YCC shorthand nomenclature names Y-DNA haplogroups and their subclades with the first letter of the major Y-DNA haplogroup followed by a dash and the name of the defining terminal SNP.[11]
Y-DNA haplogroup nomenclature is changing over time to accommodate the increasing number of SNPs being discovered and tested, and the resulting expansion of the Y-chromosome phylogenetic tree. This change in nomenclature has resulted in inconsistent nomenclature being used in different sources.[2] This inconsistency, and increasingly cumbersome longhand nomenclature, has prompted a move toward using the simpler shorthand nomenclature.
Phylogenetic structure

- Phylogenetic tree of Y-DNA haplogroups [12]
Major Y-DNA haplogroups
Haplogroups A and B
Haplogroup A is the NRY (non-recombining Y) macrohaplogroup from which all modern paternal haplogroups descend. It is sparsely distributed in Africa, being concentrated among Khoisan populations in the southwest and Nilotic populations toward the northeast in the Nile Valley. BT is a subclade of haplogroup A, more precisely of the A1b clade (A2-T in Cruciani et al. 2011), as follows:
- Haplogroup A
- Haplogroup A00
- Haplogroup A0 (formerly also A1b)
- Haplogroup A1 (also A1a-T)
- Haplogroup A1a (M31)
- Haplogroup A1b (also A2-T; P108, V221)
- Haplogroup A1b1a1 (also A2; M14)
- Haplogroup A1b1b (also A3; M32)
- Haplogroup BT (M91, M42, M94, M139, M299)
- Haplogroup B (M60)
- Haplogroup CT
Haplogroup CT (P143)
The defining mutations separating CT (all haplogroups except for A and B) are M168 and M294. The site of origin is likely in Africa. Its age has been estimated at approximately 88,000 years old,[13][14] and more recently at around 100,000[15] or 101,000 years old.[16]
Haplogroup C (M130)
- Haplogroup C (M130, M216) Found in Asia, Oceania, and North America
- Haplogroup C1 (F3393/Z1426)
- Haplogroup C1a (CTS11043)
- Haplogroup C1a1 (M8, M105, M131) Found with low frequency in Japan
- Haplogroup C1a2 (V20) Found with low frequency in Europe, Armenians, Algeria, and Nepal
- Haplogroup C1b (F1370, Z16480)
- Haplogroup C1b1 (AM00694/K281)
- Haplogroup C1b1a (B66/Z16458)
- Haplogroup C1b1a1 (M356) Found with low frequency in South Asia, Southwest Asia, and northern China
- Haplogroup C1b1a2 (B65)
- Haplogroup C1b1a2a (B67) Found among Lebbo' people in Borneo, Indonesia
- Haplogroup C1b1a2b (F725) Found among Han Chinese (Guangdong, Hunan, and Shaanxi), Dai people (Yunnan), Murut people (Brunei), Malay people (Singapore), and Aeta people (Philippines)
- Haplogroup C1b1a3 (Z16582) Found with low frequency in Saudi Arabia and Iraq
- Haplogroup C1b1b (B68) Found among Dusun people (Brunei)
- Haplogroup C1b1a (B66/Z16458)
- Haplogroup C1b2 (C-Z16582)
- Haplogroup C1b3 (B477/Z31885)
- Haplogroup C1b3a (M38) Found in Indonesia, New Guinea, Melanesia, Micronesia, and Polynesia
- Haplogroup C1b3b (M347, P309) Found among the indigenous peoples in Australia
- Haplogroup C1b1 (AM00694/K281)
- Haplogroup C1a (CTS11043)
- Haplogroup C2 (M217, P44) Found throughout Eurasia and North America, but especially among Mongols, Kazakhs, Tungusic peoples, Paleosiberians, and Na-Dené-speaking peoples
- Haplogroup C1 (F3393/Z1426)
Haplogroup D (CTS3946)
- Haplogroup D (CTS3946)
- Haplogroup D1 (M174) Found in Japan, China (especially Tibet), the Andaman Islands
- Haplogroup D1a (CTS11577)
- Haplogroup D1a1 (Z27276, Z27283, Z29263)
- Haplogroup D1a1a (M15) Found mainly in Tibetans, Qiangic peoples, Yi, and Hmong-Mien peoples
- Haplogroup D1a1b (P99) Found mainly in Tibetans, Qiangic peoples, Naxi, and Turkic peoples
- Haplogroup D1a2 (M55, M57, M64.1, M179, P12, P37.1, P41.1 (M359.1), 12f2.2) Found mainly in Japan
- Haplogroup D1a3 (Y34637) Found in Andamanese peoples (Onge, Jarawa)
- Haplogroup D1a1 (Z27276, Z27283, Z29263)
- Haplogroup D1b (L1366, L1378, M226.2) Found in Mactan Island, Philippines
- Haplogroup D1a (CTS11577)
- Haplogroup D2 (A5580.2) Found in Nigeria, Saudi Arabia and Syria
- Haplogroup D1 (M174) Found in Japan, China (especially Tibet), the Andaman Islands
Haplogroup E (M96)
- Haplogroup E (M40, M96) Found in Africa and parts of the Middle East and Europe
- Haplogroup E1 (P147)
- Haplogroup E1a (M33, M132) formerly E1
- Haplogroup E1b (P177)
- Haplogroup E1b1 (P2, DYS391p); formerly E3
- Haplogroup E1b1a (V38)
- Haplogroup E1b1a1 (M2) Found in Africa, especially among Niger–Congo-speaking populations.; formerly E3a
- Haplogroup E1b1a2 (M329) Found in Africa, especially in Ethiopia among Omotic-speaking populations.; formerly E3*
- Haplogroup E1b1b (M215)
- Haplogroup E1b1b1 (M35) Found in Horn of Africa, North Africa, the Middle East, and Europe (especially in areas near the Mediterranean and the Balkans); formerly E3b
- Haplogroup E1b1a (V38)
- Haplogroup E1b1 (P2, DYS391p); formerly E3
- Haplogroup E2 (M75)
- Haplogroup E1 (P147)
Haplogroup F (M89)
The groups descending from haplogroup F are found in some 90% of the world's population, but almost exclusively outside of sub-Saharan Africa.
F xG,H,I,J,K is rare in modern populations and peaks in South Asia, especially Sri Lanka.[12] It also appears to have long been present in South East Asia; it has been reported at rates of 4–5% in Sulawesi and Lembata. One study, which did not comprehensively screen for other subclades of F-M89 (including some subclades of GHIJK), found that Indonesian men with the SNP P14/PF2704 (which is equivalent to M89), comprise 1.8% of men in West Timor, 1.5% of Flores 5.4% of Lembata 2.3% of Sulawesi and 0.2% in Sumatra.[17][18] F* (F xF1,F2,F3) has been reported among 10% of males in Sri Lanka and South India, 5% in Pakistan, as well as lower levels among the Tamang people (Nepal), and in Iran. F1 (P91), F2 (M427) and F3 (M481; previously F5) are all highly rare and virtually exclusive to regions/ethnic minorities in Sri Lanka, India, Nepal, South China, Thailand, Burma, and Vietnam. In such cases, however, the possibility of misidentification is considered to be relatively high and some may belong to misidentified subclades of Haplogroup GHIJK.[19]
Haplogroup G (M201)
Haplogroup G (M201) originated some 48,000 years ago and its most recent common ancestor likely lived 26,000 years ago in the Middle East. It spread to Europe with the Neolithic Revolution.
It is found in many ethnic groups in Eurasia; most common in the Caucasus, Iran, Anatolia and the Levant. Found in almost all European countries, but most common in Gagauzia, southeastern Romania, Greece, Italy, Spain, Portugal, Tyrol, and Bohemia with highest concentrations on some Mediterranean islands; uncommon in Northern Europe.[20][21]
G-M201 is also found in small numbers in northwestern China and India, Bangladesh, Pakistan, Sri Lanka, Malaysia, and North Africa.
Haplogroup H (M69)
Haplogroup H (M69) probably emerged in Southern Central Asia, South Asia or West Asia, about 48,000 years BP, and remains largely prevalent there in the forms of H1 (M69) and H3 (Z5857). Its sub-clades are also found in lower frequencies in Iran, Central Asia, across the middle-east, and the Arabian peninsula.
However, H2 (P96) is present in Europe since the Neolithic and H1a1 (M82) spread westward in the Medieval era with the migration of the Roma people.
This section needs expansion. You can help by adding to it. (September 2016) |
>Text je dostupný pod licencí Creative Commons Uveďte autora – Zachovejte licenci, případně za dalších podmínek. Podrobnosti naleznete na stránce Podmínky užití.
Text je dostupný za podmienok Creative
Commons Attribution/Share-Alike License 3.0 Unported; prípadne za ďalších
podmienok.
Podrobnejšie informácie nájdete na stránke Podmienky
použitia.

